Some posts ago, we extracted some DNA from LilBUB to run a small test which revealed that BUB seems to be distantly related to Hemingway cats. However, to sequence her whole genome we require a bigger amount of DNA so we need to re-extract it from our blood sample (to remember the principles of DNA extraction check here).
But, how much DNA do you actually need to sequence the entire genome of an organism? Ideally 1 microgram should be enough. This is equivalent 0,000001 grams, which is the 50th part of a water drop. Pretty small amount, right? But in terms of our purpose more than enough!!!
After handling the sample to our sequencing facility, Norbert (one of the technicians working there) extracted BUB’s DNA. After the process he obtained a small volume of a water solution.
But how do we know how much DNA is contained there and its purity? This is determined by the ability of the DNA to absorb light, which in scientific jargon we call Absorbance.
To measure these parameters, a small drop of our water solution is loaded on a spectrophotometer. This machine emits ultraviolet light (wavelength of 260 nm) that pass through our drop, and is received by a photodetector. The less light reaching the photodetector, the more DNA is included in our sample. This analysis revealed that our sample contains up to 4 micrograms of DNA (well done Norbert!!!).
The spectrophotometer is also capable of measuring other contaminants that could be present in our sample, derived from a not so clean extraction. These contaminants absorb different types of light (with different wavelengths). The Absorbance ratios between different types of light revealed no contamination of proteins (260/280=1,82) and organic solvents (260/230=2,1), confirming the high purity of our sample.
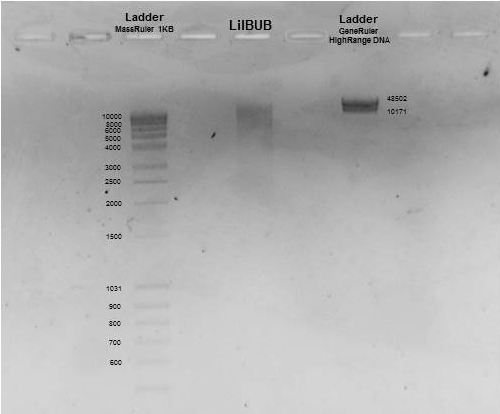
Last, we checked the size of our DNA. For that purpose, a small part the sample is loaded in an agarose gel. When we apply voltage through the gel, our DNA will migrate from top to bottom. The bigger the size of our DNA, the slower it migrates. In our picture, you can appreciate that the size of our DNA is bigger that 10 kilobases, which means that it was minimally fragmented through the whole process. Taking into account that the genetic code is composed by bases (A, C, T or G) this means than, on average, we have long DNA stretches of more than 10000 bases!!!!
Next station, preparation of a DNA library for sequencing!!!!!

2 thoughts on “A handful of BUB’s DNA”